|
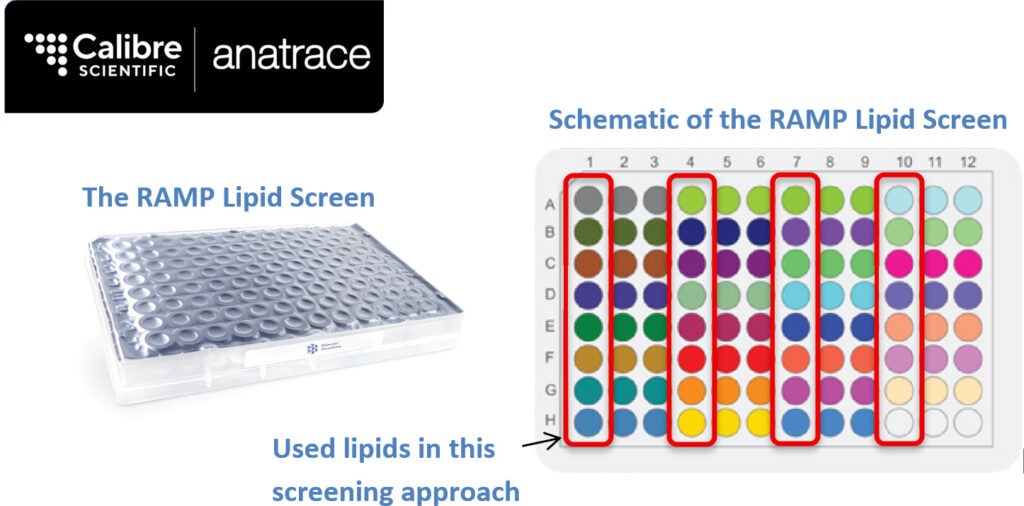
In-plate DLS Evaluation Approach of the RAMP Lipid Screen on a Transmembrane Protein ComplexThe RAMP lipid screen is a 96-well plate containing 31 lipid conditions (each in triplicate) for the high-throughput identification of lipids that stabilise membrane proteins. DLS evaluation provides maximal information about the response of an individual protein to a specific lipid while reducing the cost and reagent waste associated with lipid screening. The screen contains both individual lipids and lipid mixtures. The RAMP Lipid Screen was provided by
Each unique lipid condition is colour-coded in the schematic and is provided in triplicate. The first column of each triplicate was used in this screening approach.
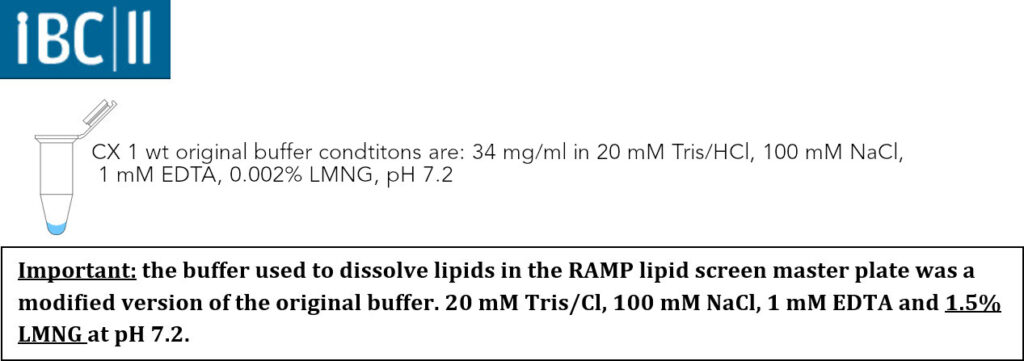
The used transmembrane portein complex was the Mitochondrial Complex I (CX 1 wt). It was contributed by the Institute of Biochemistry II of the Goethe-University of Frankfurt
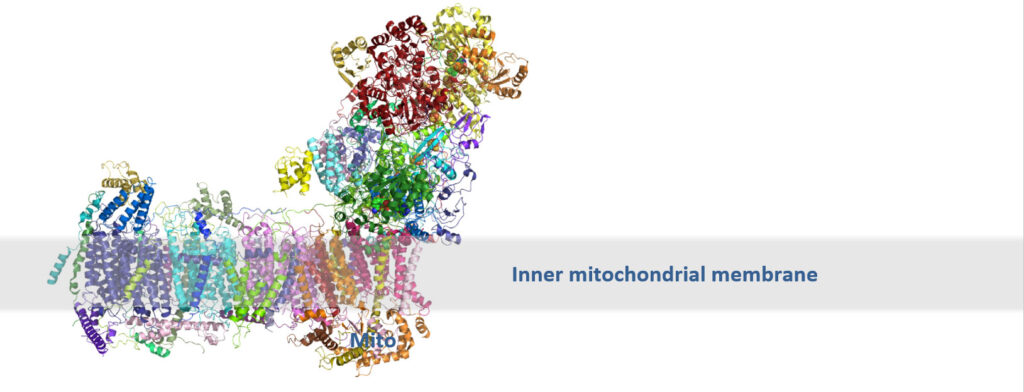
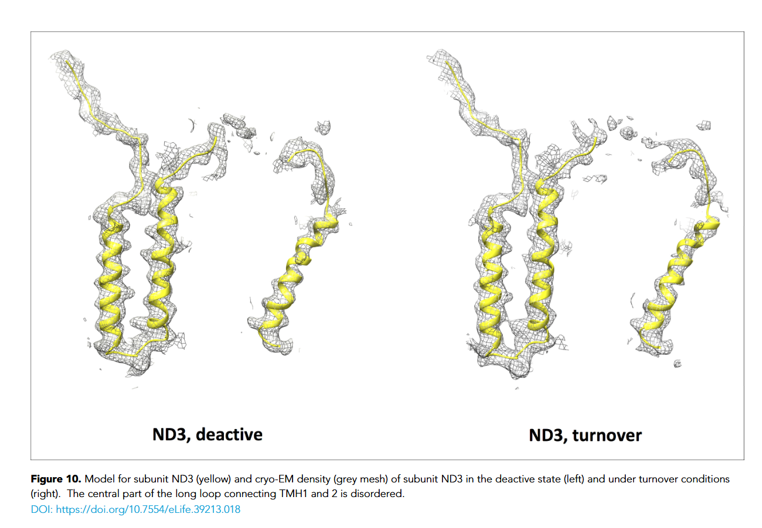
The cryo-EM structure of the Mitochondrial Complex 1 (CX 1 wt), as it is currently known The mitochondrial Complex I is a very large transmembrane protein complex (MW ~1 MDa) and is essential for aerobic energy metabolism in the cell. It utilises the redox energy released during electron transfer from NADH to ubiquinone to pump protons across the inner mitochondrial membrane. Some key positions cannot be resolved in the cryo-EM electron density of CX1wt. The change from a detergent-based buffer system to a lipid-based one may support inter-residue and backbone interactions, allowing the transmembrane region to be reconstituted and thereby stabilised. Reducing random orientations is expected to increase electron density. 3D single-particle cryo-EM on the CX 1 wt showed low occupancy of the multi-α-helical transmembrane region. The aim of this screen is to improve this
A major challenge for this complex is the low occupancy of the transmembrane α-helices due to randomisation of their relative orientation. The aim of the screen is to replace LMNG with a lipid that maintains the protein in a stable, soluble state without inducing aggregation or precipitation.
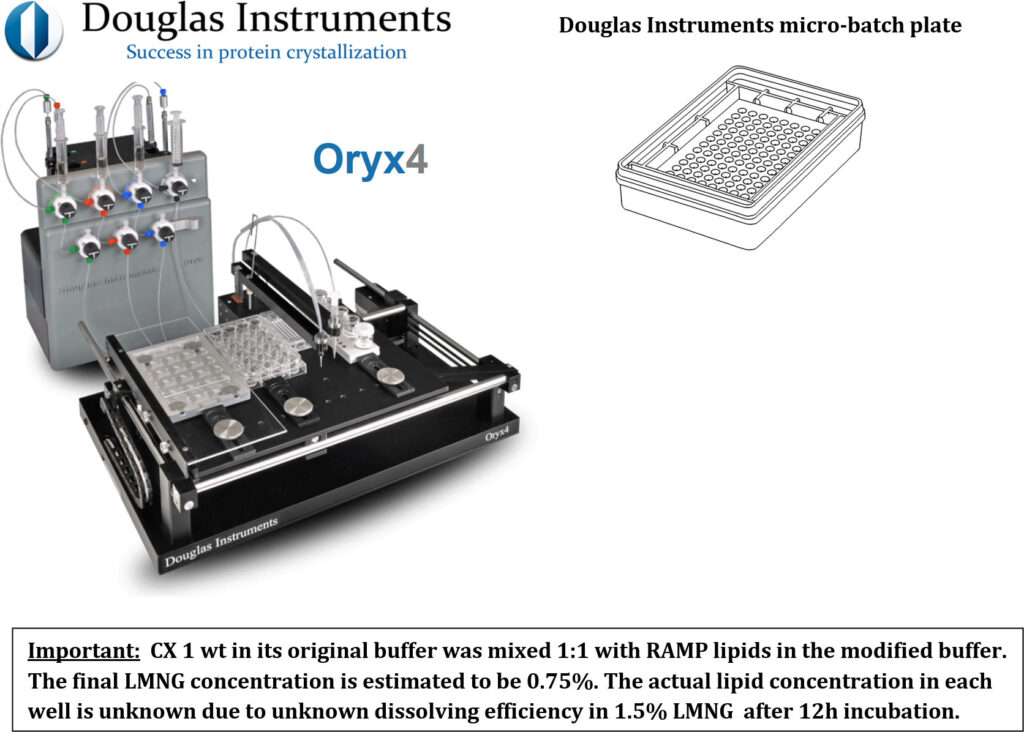
The Oryx4 Protein Crystallization Robot used for the RAMP lipid screen in micro-batch plates
In-plate DLS data collection and evaluation were carried out by
DLS measurement parameters:Duration: 10 s Number of measurements: 10 Number of scans: 10 Scan delay time = 120 min. Measurement temperature 20 °C Laser intensity 10% Sample Volume 0.2 µl Paraffin oil seal pre-incubation for 30 min.
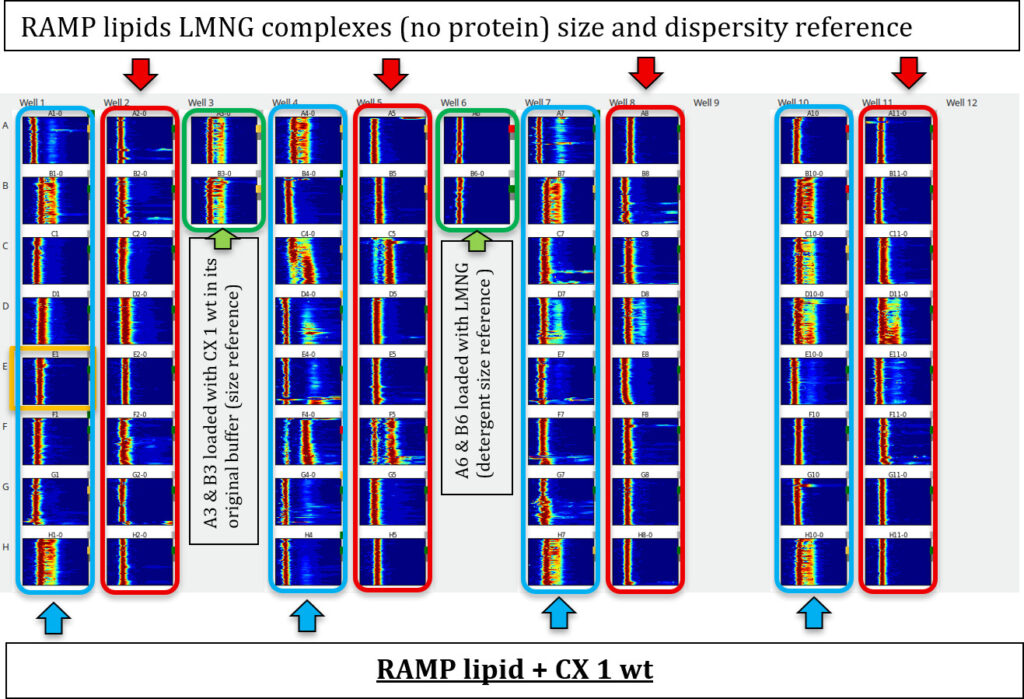
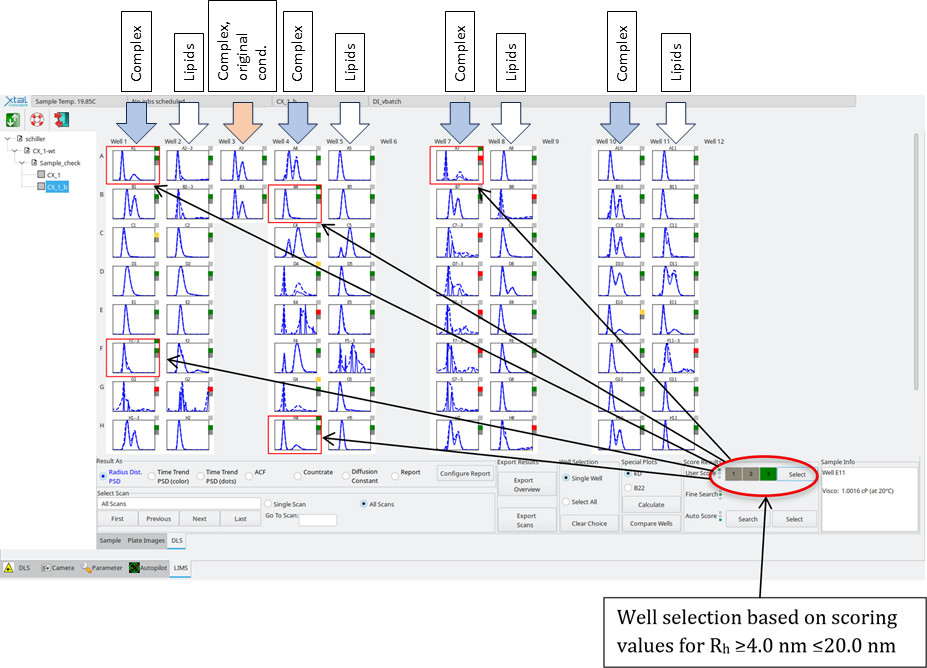
Screening ResultsImage overview of the RAMP lipid Screen/CX 1 wt (200 - 500 nl/well)
Slight precipitation was observed in some droplets; however, most droplets were clear. Because the lipids were solubilised by LMNG, they generate a strong DLS signal on their own, the DLS evaluation followed a characteristic size-distribution signature. For comparison, the LMNG lipid buffer was loaded without protein to determine the resulting size distribution. The same condition was then loaded with protein to observe any size differences. The criteria were based on changes in particle size, distribution profile, and overall monodispersity relative to the lipid-only control. The evaluation was based on the following principle: Lipid LNMG complex ≠ Protein lipid complex ~ Protein in original buffer condition
RAMP lipid Screen DLS result on CX 1 wt, solubility and long term stability (day 1 - 11) after loading
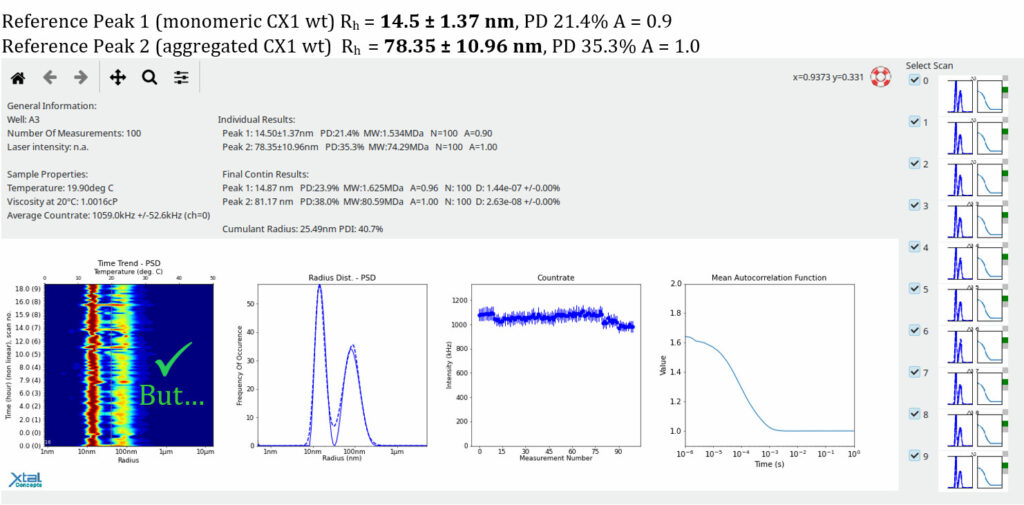
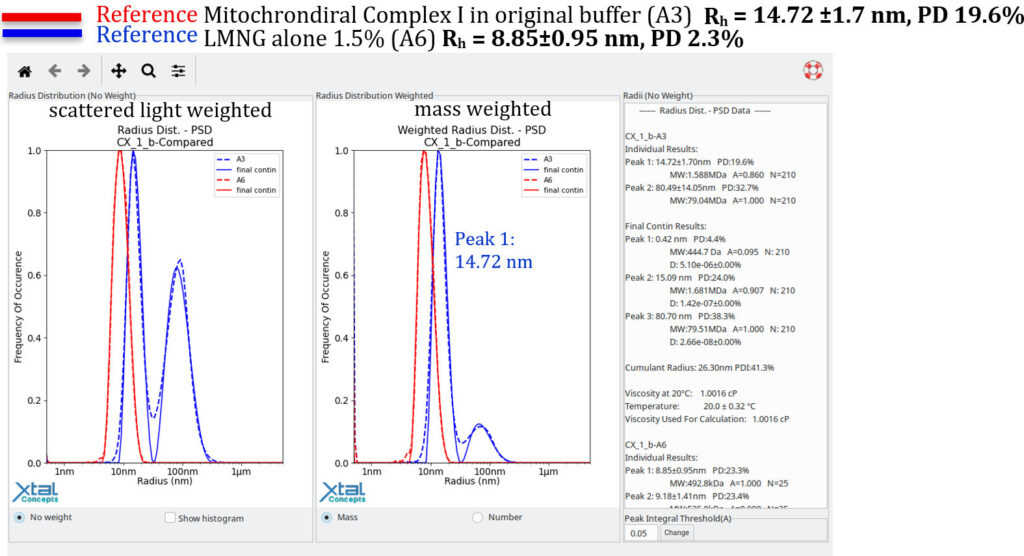
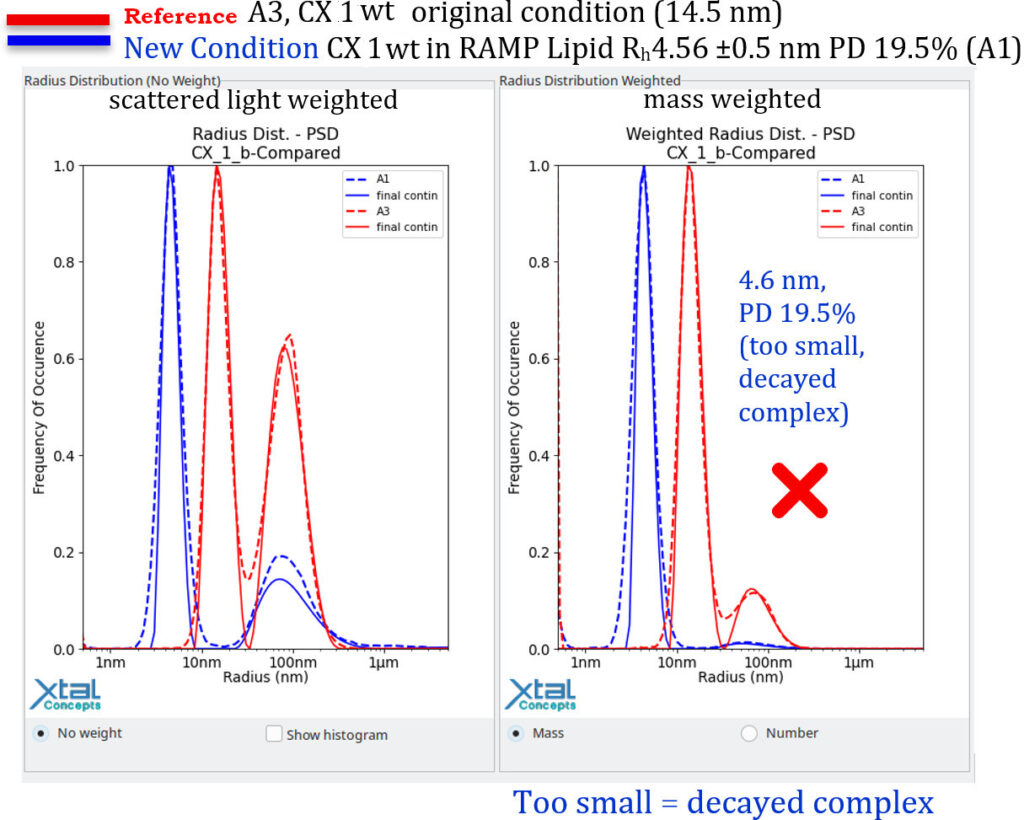
RAMP lipid Screen DLS EvaluationDLS results of the 0riginal condition of the Mitochondrial Complex 1 (34 mg/ml, Well A3), 15h observationThe following plot, presented in report format, shows the original condition and is used as a reference for evaluating new lipid conditions. A newly identified condition should exhibit particles of similar size to the reference condition while clearly differing from lipid alone (i.e., lipid–LMNG complexes). As an additional criterion, the size distribution should trend towards increased monodispersity.
The goal is to achieve the same size distribution, or a more mono disperse distribution, with the same absolute size of the mitochondrial complex I, but in a lipid condition.
RAMP lipid Screen/CX 1 wt, Identification of the most suited Lipids by Scoring:What are the improved lipid conditions now? We checked two criteria: A: The protein-lipid complex must differ from the dissolved lipid. B: The sample should be similar in size to the complex in the original buffer condition and ideally less poly disperse. We checked each well pair by an applying an auto-scoring algorithm (green square in the top right-hand corner) if these criteria were met. The highest score (green) was applied to such wells, which were then selected based on the following values: Rh ≥ 4.0 nm ≤ 20.0 nm.
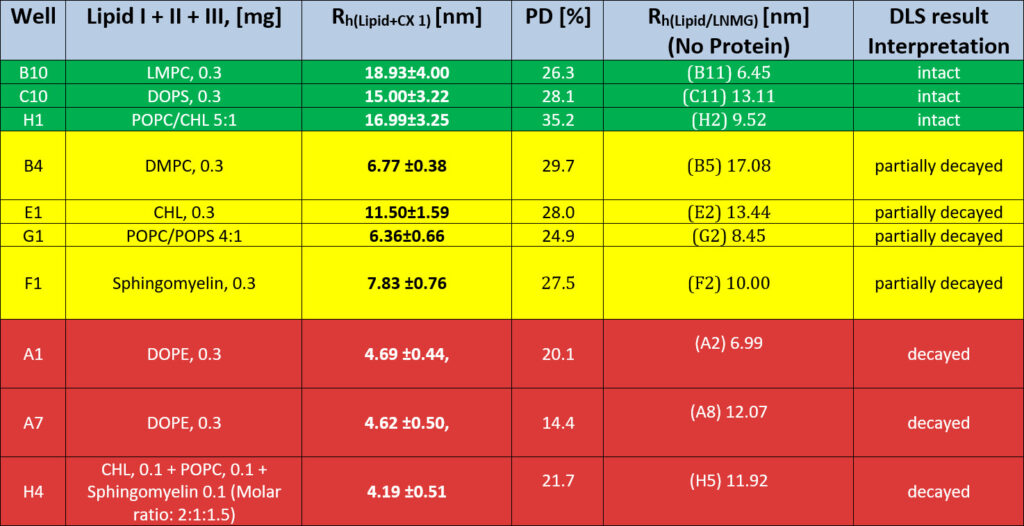
After repetitive scoring, a set of newly identified conditions emerged. However, some of these may represent intermediate or partially stabilising states rather than fully optimal lipid environments. Table of Summarized RAMP lipid Screen/CX 1 wt, Scoring ResultsAfter repetitive scoring, a set of newly found conditions emerges. However, some of these could be interpreted as partially or fully decayed mitochondrial complex 1.
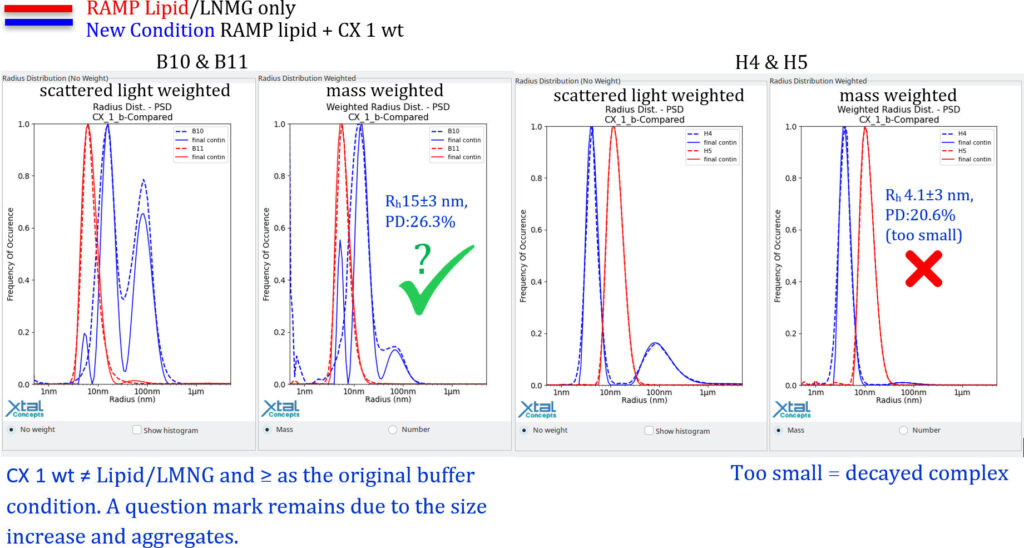
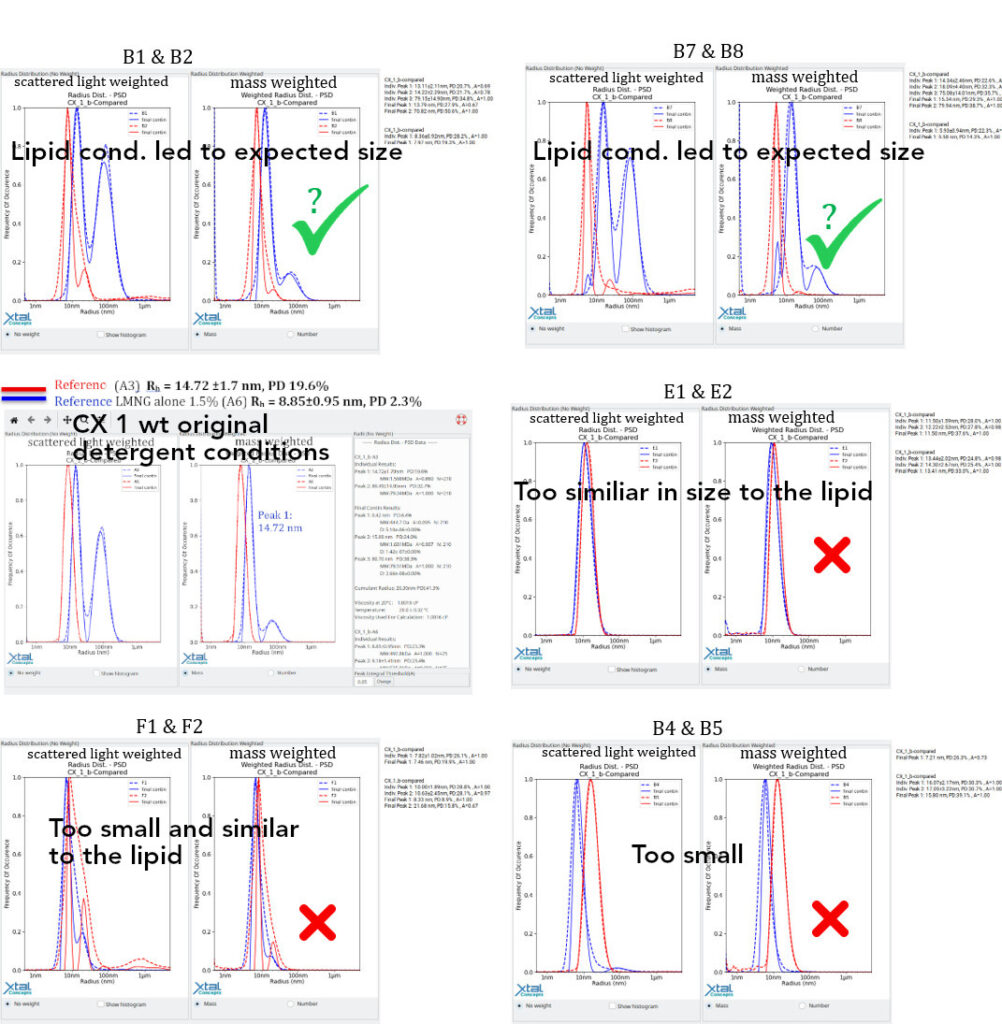
Individual Well DLS Evaluation of the RAMP lipid screen by Size ComparisonWhether a RAMP lipid improves the solubility by detergent exchange can be revealed by comparison between wells, of RAMP lipid only with RAMP lipid + CX 1 wt.
CX 1 wt, Detergent (LMNG) Comparison
Decayed CX 1 wt by Comparison with the original conditions
The general questions are: What happens to the lipid LMNG complex in the presence of the protein? Has it been fully adsorbed by the complex? In that case, is CX 1 wt acting as an emulsifying agent? See also B4 & B5. Lipid conditions for single particle Cryo-EM samples (intact lipid CX 1 wt complex?):Although improved monodispersity could not be achieved through LMNG lipid exchange, we expect the lipid conditions to be suitable for single-particle 3D cryoEM imaging of the complex. As the original conditions were similar in terms of size distribution and had already been successfully imaged using single-particle 3D cryo-EM, it is likely that there is a sufficient quantity of the complex for this type of investigation.
A question mark remains due to result is not confirmed yet by Cryo-EM
These experiments would not have been possible without the support of four contributing parties:
I would like to express my sincere gratitude to all of you for your close cooperation and the trust you have placed in the project and in us. Thank you for your interest in this project! Kind regards, May 24, 2026 |
|
; |

|
|
|